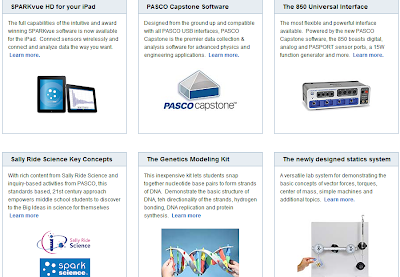
Using the Ideal Gas equation, we measured the number of moles: From the Pasco Capstone software, we obtained the values of pressure and temperature. This pressure was measured by special Pasco detectors. We measured the volume of the collected gas, then we applied pressure by pushing down the plunge. We collected gas sample from our cultures grown in anaerobic conditions into the special measuring cylinder. Below, you can see the photo of our measurement setup:
#Pasco capstone software software#
The Ideal Gas Apparatus, PASCO sensor PASCO Capstone software were used to measure pressure and temperature of the system. The pressure and temperature of samples were measured on 22.10.19. They were flushed by nitrogen gas in order to make anaerobic conditions. There were two cultures cultured in two different times (04.10.19 and 16.10.19). This leads us to be sure that hydrogen is being produced: Stopper was punctured using need and excess gas was let out straight to the flame of match instead of usual “Pop” sound we heard whistle in both cases and same reaction of flame to the gas that went out in control and our culture tube. Zinc + HCl reaction was done in the same sealed anaerobic tube. Pop test was performed as a classical way to detect hydrogen in the air one of our tubes showed exact same result to a lesser extent as the control reaction of Zinc + Hydrochloric Acid. This is supported by this survival assay and thus hydrogen presence by HydA activity. 5 days later tubes were assessed on paper colour to confirm survival.Īs Ducat stated limited growth is supported in Cyanobacteria as they use Hydrogen as reductant in alternative metabolic pathways. Tubes were flushed with nitrogen until air is replaced. It was soaked in the medium with sulfide and w/o sulfide. From Figure 7, it can be seen that transformed HydA+ cells differ significantly from the wild-type by its color illustrating the higher survival rate compared to the wild-type strains.īlotted paper was also assessed using survival assay with the same condition only few adjustments for the paper. The samples of nearly same OD were constantly shaken in the shaking incubator.
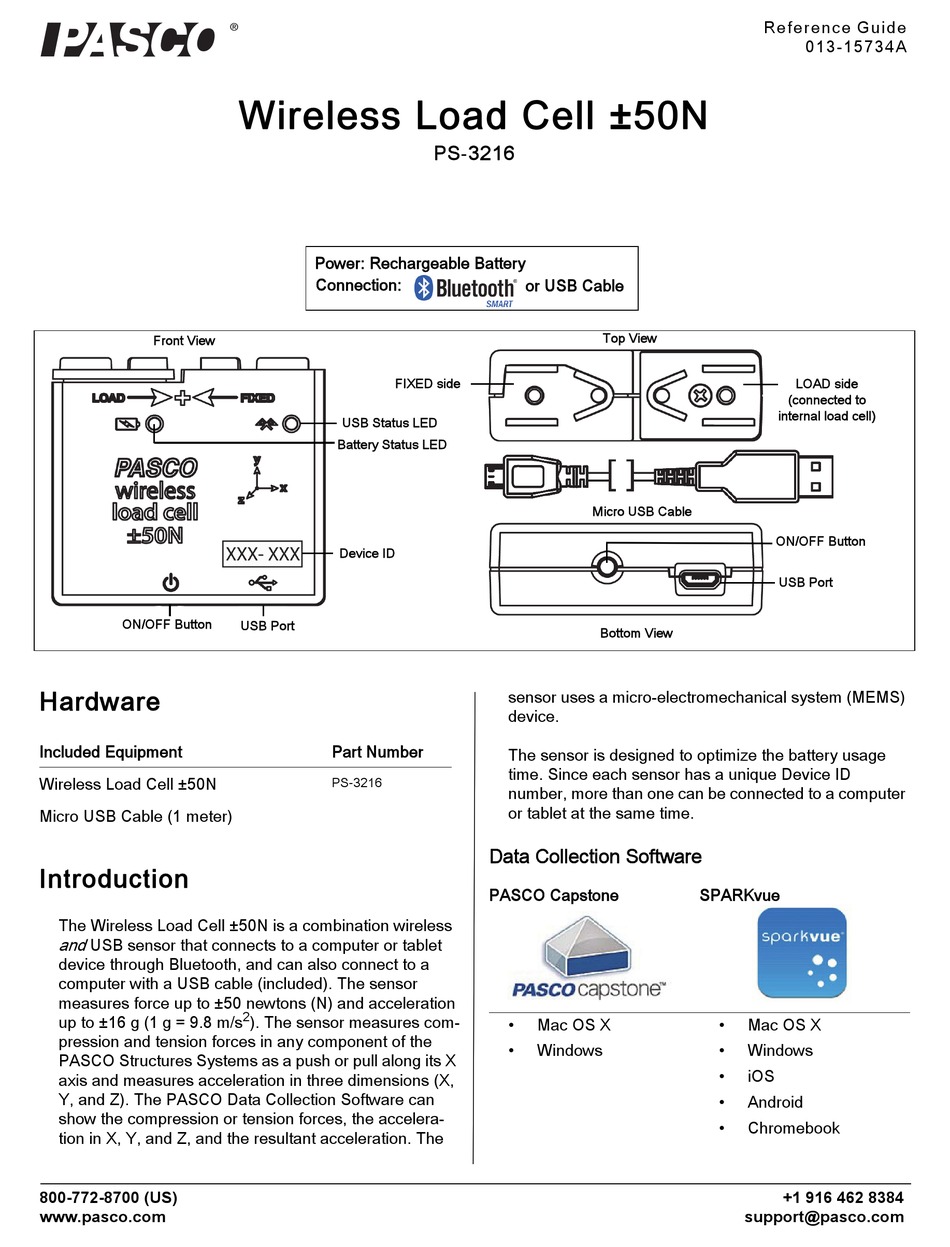
The effect of genetic modification on survivability of cyanobacteria in a hydrogen sulfide (H2S) rich and anaerobic environment was assessed qualitatively. HydA+PsbA1 amplification from the construct integrated into cyanobacteria Out of five samples, the 5th band without ladder had an expected band for ~3.1 kb.įigure6. NSb primer was used due to complications during amplification process with HydA primers.

To verify the integration of construct into cyanobacteria psbA1+hydA(~3.1 kb) amplification was performed using forward psbA1 and reverse NSb primers. As a negative control p233 (also known as MF) without insert was used which did not show any band for HydA as expected.įigure5. Overall, four samples were assessed for the HydA and three were positive for the gene as bands corresponding to ~1.7-1.8 kb were visualized on the gel. To ensure the presence of HydA gene in the construct, the samples were linearized by EcoRV and used as template for HydA amplification (see Figure 5). Linear assembled construct digested by BamH gel results All samples were linearized with BamHI-HF beforehand.įigure4. HydA and PsbA1 assembly gel resultsĪfter transformation of the assembled construct into the DH5alpha cloning strain, DNA extraction results illustrated bands higher than empty vector p233 (also known as MF (Maturation Factor)). As it can be seen from Figure 1 and Figure 2 successful amplification of genes of interest was verified via gel electrophoresis.įigure1: PsbA1 amplification gel results.Īmplified HydA and psbA1 genes were assembled via Polymerase Cycling Assembly (PCA) and also verified via gel electrophoresis as shown on Figure 3.įigure 3. To assemble psbA1 and HydA genes, firstly, amplification of psbA1 (503 bp) from p233(11830 bp) and HydA(1998 bp) from plasmid PAM (5004 bp) were done. Plasmid p233 that contains hydrogenase maturation factors HydEF and HydG under psbA1 promoter and HydA under psbA1 promoter in 0015 plasmids. Due to complications with amplification and assembly of synthetic sequences of genes that we ordered like complex stable secondary structures and a lot of long repeats we switched to using plasmids that we received from as a kind gesture from Daniel C.
